Publications
A current list of my publications is available on my Google Scholar profile.
My full CV is also available for download here.
First Author Publications
Weisberg A.J., Rahman A., Backus D., Tyavanagimatt P., Chang J.H., and Sachs J.L. (2022) Pangenome evolution reconciles robustness and instability of rhizobial symbiosis. mBio ahead of print. https://doi.org/10.1128/mbio.00074-22
Weisberg A.J., Miller M., Ream W., Grünwald N., & Chang J.H. (2021). Diversification of plasmids in a genus of pathogenic and nitrogen-fixing bacteria. Philosophical Transactions of the Royal Society B, 377(1842), p20200466 https://doi.org/10.1098/rstb.2020.0466
Weisberg, A. J., Grünwald, N. J., Savory, E. A., Putnam, M. L., & Chang, J. H. (2021). Genomic approaches to plant-pathogen epidemiology and diagnostics. Annual Review of Phytopathology, 59. https://doi.org/10.1146/annurev-phyto-020620-121736
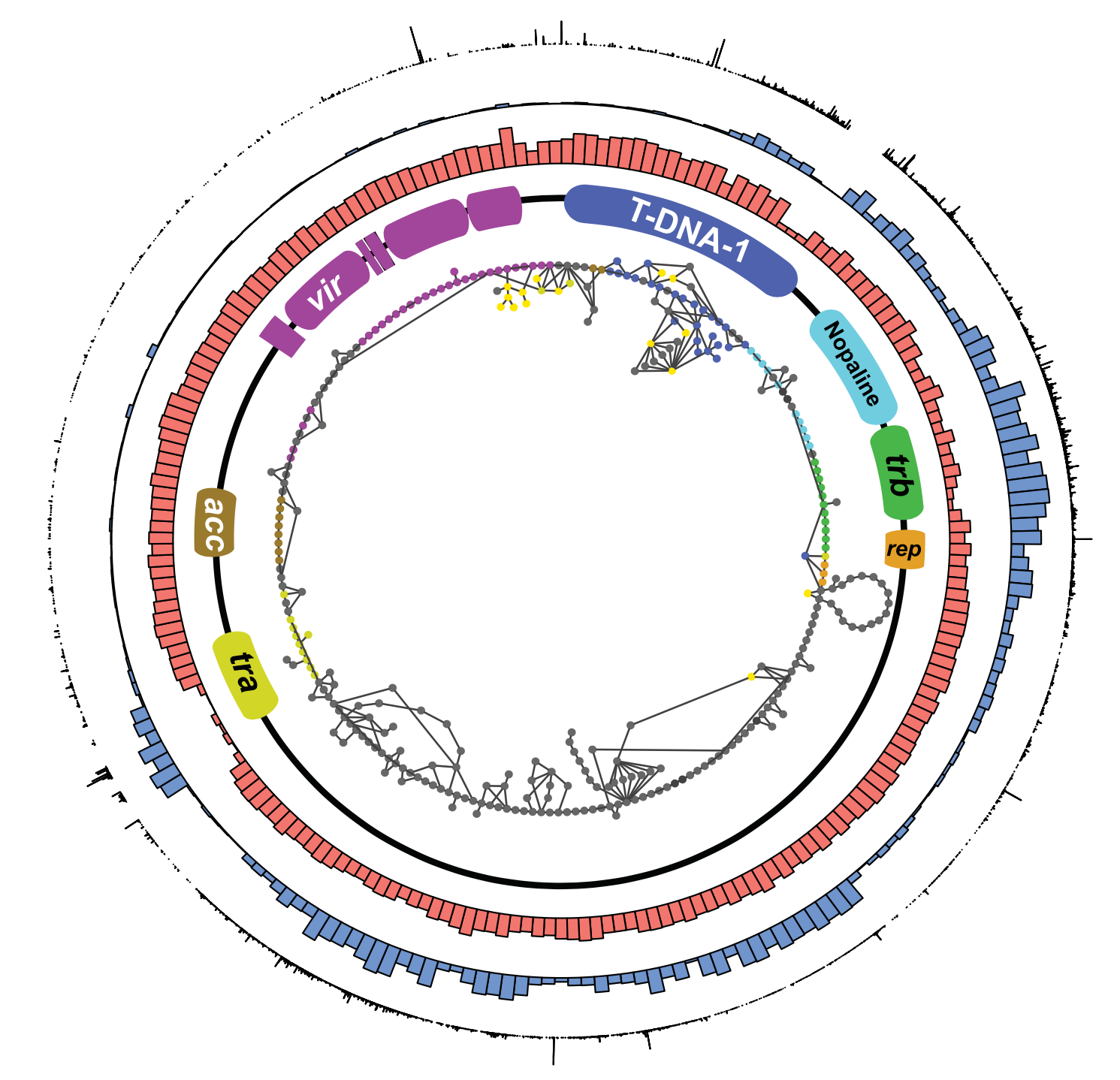
Weisberg, A. J., Davis, E. W., Tabima, J., Belcher, M. S., Miller, M., Kuo, C.-H., Loper, J. E., Grünwald, N. J., Putnam, M. L., & Chang, J. H. (2020). Unexpected conservation and global transmission of agrobacterial virulence plasmids. Science, 368(6495). https://doi.org/10.1126/science.aba5256
Savory, E. A.*, Weisberg, A. J.*, Stevens, D. M., Creason, A. L., Fuller, S. L., Pearce, E. M., & Chang, J. H. (2020). Phytopathogenic rhodococcus have diverse plasmids with few conserved virulence functions. Frontiers in Microbiology, 11, 1022. https://doi.org/10.3389/fmicb.2020.01022
Weisberg, A. J., Kramer, C. G., Kotha, R. R., Luthria, D. L., Chang, J. H., & Clarke, C. R. (2020). A novel species-level group of streptomyces exhibits variation in phytopathogenicity despite conservation of virulence loci. Molecular Plant-Microbe Interactions, MPMI–06. https://doi.org/10.1094/MPMI-06-20-0164-R
Savory, E. A.*, Fuller, S. L.*, Weisberg, A. J.*, Thomas, W. J., Gordon, M. I., Stevens, D. M., Creason, A. L., Belcher, M. S., Serdani, M., Wiseman, M. S., Grünwald, N. J., Putnam, M. L., & Chang, J. H. (2017). Evolutionary transitions between beneficial and phytopathogenic rhodococcus challenge disease management. eLife, 6, e30925. https://doi.org/10.7554/elife.30925
Weisberg, A. J., Kim, G., Westwood, J. H., & Jelesko, J. G. (2017). Sequencing and de novo assembly of the toxicodendron radicans (poison ivy) transcriptome. Genes, 8(11), 317. https://doi.org/10.3390/genes8110317
Davis II, E. W.*, Weisberg, A. J.*, Tabima, J. F., Grünwald, N. J., & Chang, J. H. (2016). Gall-ID: Tools for genotyping gall-causing phytopathogenic bacteria. PeerJ, 4, e2222. https://doi.org/10.7717/peerj.2222
Weisberg, A. J., Elmarakeby, H. A., Heath, L. S., & Vinatzer, B. A. (2015). Similarity-based codes sequentially assigned to ebolavirus genomes are informative of species membership, associated outbreaks, and transmission chains. Open Forum Infectious Diseases, 2, ofv024. https://doi.org/10.1093/ofid/ofv024
Co-Author Publications
Chou L., Lin Y.C., Haryono M., Santos M.N.M., Cho S.T., Weisberg A.J., Wu C.F., Chang J.H., Lai E.M., and Kuo C.H. (2022). Modular evolution of secretion systems and virulence plasmids in a bacterial species complex. BMC biology 20 article 16. https://doi.org/10.1186/s12915-021-01221-y
Nguyen H.P., Weisberg A.J., Chang J.H., and Clarke C.R.. Streptomyces caniscabiei sp. nov., which causes potato common scab and is distributed across the world. International Journal of Systematic and Evolutionary Microbiology 72 (1), 005225 https://doi.org/10.1099/ijsem.0.005225
Bland R.N., Johnson J.D., Waite-Cusic J.G., Weisberg A.J., Riutta E.R., Chang J.H., Kovacevic J. (2021). Application of whole genome sequencing to understand diversity and presence of genes associated with sanitizer tolerance in Listeria monocytogenes from produce handling sources. Foods 10.10 p. 2454. https://doi.org/10.3390/foods10102454
Wu, C.-F., Weisberg, A. J., Davis, E. W., Chou, L., Khan, S., Lai, E.-M., Kuo, C.-H., & Chang, J. H. (2021). Diversification of the type VI secretion system in agrobacteria. mBio, 12(5), e01927–21. https://doi.org/10.1128/mBio.01927-21
Quides, K. W., Weisberg, A. J., Trinh, J., Salaheldine, F., Cardenas, P., Lee, H.-H., Jariwala, R., Chang, J. H., & Sachs, J. L. (2021). Experimental evolution can enhance benefits of rhizobia to novel legume hosts. Proceedings of the Royal Society B, 288(1951), 20210812. https://doi.org/10.1098/rspb.2021.0812
Torres-Martı́nez, L., Porter, S. S., Wendlandt, C., Purcell, J., Ortiz-Barbosa, G., Rothschild, J., Lampe, M., Warisha, F., Le, T., Weisberg, A. J., Chang, J. H., & Sachs, J. L. (2021). Evolution of specialization in a plant-microbial mutualism is explained by the oscillation theory of speciation. Evolution, 75(5), 1070–1086. https://doi.org/10.1111/evo.14222
Nofiani, R., Weisberg, A. J., Tsunoda, T., Panjaitan, R. G. P., Brilliantoro, R., Chang, J. H., Philmus, B., & Mahmud, T. (2020). Antibacterial potential of secondary metabolites from Indonesian marine bacterial symbionts. International Journal of Microbiology, 2020. https://doi.org/10.1155/2020/8898631
Turner, S. E., Pang, Y.-Y., O’Malley, M. R., Weisberg, A. J., Fraser, V. N., Yan, Q., Chang, J. H., & Anderson, J. C. (2020). A deoR-type transcription regulator is required for sugar-induced expression of Type III secretion-encoding genes in pseudomonas syringae pv. Tomato DC3000. Molecular Plant-Microbe Interactions, 33(3), 509–518. https://doi.org/10.1094/mpmi-10-19-0290-r
Vasebi, Y., Khakvar, R., Tian, L., Moubarak, P., Valentini, F., Weisberg, A. J., & Vinatzer, B. A. (2020). Phenotypic characterization and phylogenetic analysis of pseudomonas syringae strains associated with canker disease on apricot in iran within the context of the global genetic diversity of the p. Syringae complex. European Journal of Plant Pathology, 158, 545–560. https://doi.org/10.1007/s10658-020-02101-x
Zhou, W., Posri, P., Abugrain, M. E., Weisberg, A. J., Chang, J. H., & Mahmud, T. (2020). Biosynthesis of the nuclear factor of activated t cells inhibitor NFAT-133 in streptomyces pactum. ACS Chemical Biology. https://doi.org/10.1021/acschembio.0c00775
Gano-Cohen, K. A., Wendlandt, C. E., Al Moussawi, K., Stokes, P. J., Quides, K. W., Weisberg, A. J., Chang, J. H., & Sachs, J. L. (2020). Recurrent mutualism breakdown events in a legume rhizobia metapopulation. Proceedings of the Royal Society B, 287(1919),
Lopes, L. D., Weisberg, A. J., Davis, E. W., Varize, C. de S., Pereira e Silva, M. de C., Chang, J. H., Loper, J. E., & Andreote, F. D. (2019). Genomic and metabolic differences between pseudomonas putida populations inhabiting sugarcane rhizosphere or bulk soil. PLOS One, 14(10), e0223269. https://doi.org/10.1371/journal.pone.0223269
O’Malley, M. R., Weisberg, A. J., Chang, J. H., & Anderson, J. C. (2019). Re-evaluation of a Tn 5:: gacA mutant of pseudomonas syringae pv. Tomato DC3000 uncovers roles for uvrC and anmK in promoting virulence. PLOS One, 14(10), e0223637. https://doi.org/10.1371/journal.pone.0223637
Thapa, S. P., Davis, E. W., Lyu, Q., Weisberg, A. J., Stevens, D. M., Clarke, C. R., Coaker, G., & Chang, J. H. (2019). The evolution, ecology, and mechanisms of infection by gram-positive, plant-associated bacteria. Annual Review of Phytopathology, 57, 341–365. https://doi.org/10.1146/annurev-phyto-082718-100124
Chang, J. H., Putnam, M. L., Grünwald, N. J., Savory, E. A., Fuller, S. L., & Weisberg, A. J. (2018). Response to comments on “evolutionary transitions between beneficial and phytopathogenic rhodococcus challenge disease management”. eLife, 7, e35852. https://doi.org/10.7554/eLife.35852
Davis, E. W., Tabima, J. F., Weisberg, A. J., Lopes, L. D., Wiseman, M. S., Wiseman, M. S., Pupko, T., Belcher, M. S., Sechler, A. J., Tancos, M. A., Schroeder, B. K., Murray, T. D., Luster, D. G., Schneider, W. L., Rogers, E. E., Andreote, F. D., Grünwald, N. J., Putnam, M. L., & Chang, J. H. (2018). Evolution of the US biological select agent rathayibacter toxicus. MBio, 9(4). https://doi.org/10.1128/mbio.01280-18
Dickinson, C. C., Weisberg, A. J., & Jelesko, J. G. (2018). Transient heterologous gene expression methods for poison ivy leaf and cotyledon tissues. HortScience, 53(2), 242–246. https://doi.org/10.21273/hortsci12421-17
Eckshtain-Levi, N., Weisberg, A. J., & Vinatzer, B. A. (2018). The population genetic test Tajima’s D identifies genes encoding pathogen-associated molecular patterns and other virulence-related genes in ralstonia solanacearum. Molecular Plant Pathology, 19(9), 2187–2192. https://doi.org/10.1111/mpp.12688
Lopes, L. D., Davis, E. W., Pereira e Silva, M. de C., Weisberg, A. J., Bresciani, L., Chang, J. H., Loper, J. E., & Andreote, F. D. (2018). Tropical soils are a reservoir for fluorescent pseudomonas spp. biodiversity. Environmental Microbiology, 20(1), 62–74. https://doi.org/10.1111/1462-2920.13957
Lopes, L. D., Pereira e Silva, M. de C., Weisberg, A. J., Davis, E. W., Yan, Q., Varize, C. de S., Wu, C.-F., Chang, J. H., Loper, J. E., & Andreote, F. D. (2018). Genome variations between rhizosphere and bulk soil ecotypes of a pseudomonas koreensis population. Environmental Microbiology, 20(12), 4401–4414. https://doi.org/10.1111/1462-2920.14363
Muchero, W., Sondreli, K. L., Chen, J.-G., Urbanowicz, B. R., Zhang, J., Singan, V., Yang, Y., Brueggeman, R. S., Franco-Coronado, J., Abraham, N., Yang, J.-Y., Moremen, K. W., Weisberg, A. J., Chang, J. H., Lindquist, E., Barry, K., Ranjan, P., Jawdy, S., Schmutz, J., … LeBoldus, J. M. (2018). Association mapping, transcriptomics, and transient expression identify candidate genes mediating plant-pathogen interactions in a tree. Proceedings of the National Academy of Sciences, 115(45), 11573–11578. https://doi.org/10.1073/pnas.1804428115
Fuller, S. L., Savory, E. A., Weisberg, A. J., Buser, J. Z., Gordon, M. I., Putnam, M. L., & Chang, J. H. (2017). Isothermal amplification and lateral-flow assay for detecting crown-gall-causing Agrobacterium spp. Phytopathology, 107(9), 1062–1068. https://doi.org/10.1094/phyto-04-17-0144-r
Moore, E. R., Bullington, B. S., Weisberg, A. J., Jiang, Y., Chang, J., & Halsey, K. H. (2017). Morphological and transcriptomic evidence for ammonium induction of sexual reproduction in thalassiosira pseudonana and other centric diatoms. PLOS One, 12(7), e0181098. https://doi.org/10.1371/journal.pone.0181098
Vinatzer, B. A., Weisberg, A. J., Monteil, C. L., Elmarakeby, H. A., Sheppard, S. K., & Heath, L. S. (2017). A proposal for a genome similarity-based taxonomy for plant-pathogenic bacteria that is sufficiently precise to reflect phylogeny, host range, and outbreak affiliation applied to pseudomonas syringae sensu lato as a proof of concept. Phytopathology, 107(1), 18–28. https://doi.org/10.1094/phyto-07-16-0252-r
Tabima, J. F., Everhart, S. E., Larsen, M. M., Weisberg, A. J., Kamvar, Z. N., Tancos, M. A., Smart, C. D., Chang, J. H., & Grünwald, N. J. (2016). Microbe-ID: An open source toolbox for microbial genotyping and species identification. PeerJ, 4, e2279. https://doi.org/10.7717/peerj.2279
Clarke, C. R., Studholme, D. J., Hayes, B., Runde, B., Weisberg, A., Cai, R., Wroblewski, T., Daunay, M.-C., Wicker, E., Castillo, J. A., & Vinatzer, B. A. (2015). Genome-enabled phylogeographic investigation of the quarantine pathogen ralstonia solanacearum race 3 biovar 2 and screening for sources of resistance against its core effectors. Phytopathology, 105(5), 597–607. https://doi.org/10.1094/phyto-12-14-0373-r
Dornfeld, C., Weisberg, A. J., KC, R., Dudareva, N., Jelesko, J. G., & Maeda, H. A. (2014). Phylobiochemical characterization of class-Ib aspartate/prephenate aminotransferases reveals evolution of the plant arogenate phenylalanine pathway. Plant Cell, 26(7), 3101–3114. https://doi.org/10.1105/tpc.114.127407
Manuscripts in press or in preparation
”Dynamic interactions between mega-sized symbiosis integrative and conjugative elements and bacterial chromosomes” Weisberg A.J., Sachs J.L., Chang J.H. under review.
”Dual adhesive unipolar polysaccharides synthesized by overlapping biosynthetic pathways in Agrobacterium tumefaciens” Onyeziri M.C., Natarajan R., Hardy G.G., Xu J., Reynolds I.P., Kim J., Merritt P.M., Danhorn T., Hibbing M.E., Weisberg A.J., Chang J.H., Fuqua C. Preprint available at https://www.biorxiv.org/content/10.1101/2021.04.22.440995v1
Plus three additional manuscripts in preparation.